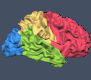
Exhibition at Annual OHBM Meeting in Seoul, South Korea
20 06, 24
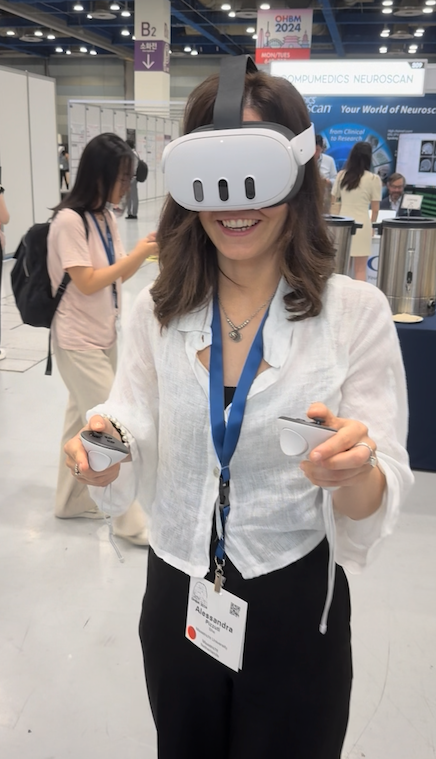
If you are attending the Human Brain Mapping conference in Seoul, South Korea, please visit our booth (#209) and you'll meet BrainVoyager experts answering your questions! We are thrilled to show you several new applications and updates of existing software, including BrainVoyager 23.2 with its amazing new volume renderer (see figure below). We also show the near-release version of the desktop-quality software BrainViewer running on almost every operating system with a touch-driven user interface on mobile platforms including iOS and Android. Like the recently released Brain Tutor 3D app, BrainViewer not only runs on all major desktop and mobile platforms but it also comes as a WebApp (based on WebAssembly) allowing to launch the app from most web browsers without installing anything. The BrainViewer app not only supports BrainVoyager files but also NIfTI and GIfTI files in a powerful app that can be used to visualise volume and surface datasets with overlaid regions, maps and invoked time courses. The app is so general that it can be used by BrainVoyager users as well as users of other software packages to inspect datasets and to create beautiful publication-ready figures and movies. BrainViewer also runs on VR headsets - come to our booth and try out Brain Viewer XR using a Meta Quest 3 headset!

We also highlight our latest open source contributions to the software LayNII for state-of-the-art laminar and columnar fMRI analysis. We also present Turbo-BrainVoyager 4.4 that now supports multi-slice DICOM files (used increasingly by SIEMENS) and NIfTI files (used by PHILIPS). TBV 4.4 also comes with a new real-time data simulator that is also included in the Turbo-BrainVoyager 4.4 EDU version. Our EDU products are the ideal tools to teach (e.g. in courses) and (self-)learn how to analyse and visualise structural and functional neuroimaging data from basic to advanced tools including scripting and programming using Python. In our partnership with NIRx, we are excited to present the latest version of our real-time fNIRS software Turbo-Satori and a major new version of our fNIRS offline software Satori that now integrates calls into Python tools, for example, in graphically designed data processing workflows.

We also highlight our latest open source contributions to the software LayNII for state-of-the-art laminar and columnar fMRI analysis. We also present Turbo-BrainVoyager 4.4 that now supports multi-slice DICOM files (used increasingly by SIEMENS) and NIfTI files (used by PHILIPS). TBV 4.4 also comes with a new real-time data simulator that is also included in the Turbo-BrainVoyager 4.4 EDU version. Our EDU products are the ideal tools to teach (e.g. in courses) and (self-)learn how to analyse and visualise structural and functional neuroimaging data from basic to advanced tools including scripting and programming using Python. In our partnership with NIRx, we are excited to present the latest version of our real-time fNIRS software Turbo-Satori and a major new version of our fNIRS offline software Satori that now integrates calls into Python tools, for example, in graphically designed data processing workflows.
BrainVoyager 23 Now Available with Native Apple Silicon Support
17 10, 23
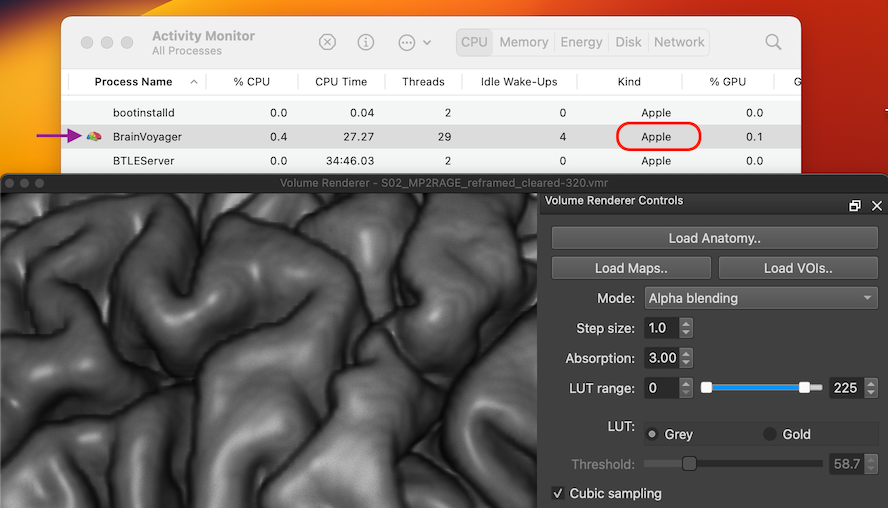
We are happy to announce the availability of BrainVoyager 23.0 and BrainVoyager 23.0 EDU! This is a major release introducing important new features. A major focus of this release was maximising the high-performance, energy-efficient capabilities of Apple Silicon Macs for BrainVoyager. Every aspect of BrainVoyager has been considered with respect to the question how to benefit best from a native Apple Silicon implementation. The result is amazing graphics and compute performance based on a native arm64 compiled binary and usage of Apple's Metal and Accelerate frameworks. The snapshot below shows an entry of BrainVoyager 23 in Activity Monitor: the "Apple" entry in the "Kind" column identifies BrainVoyager as a native Apple Silicon application.
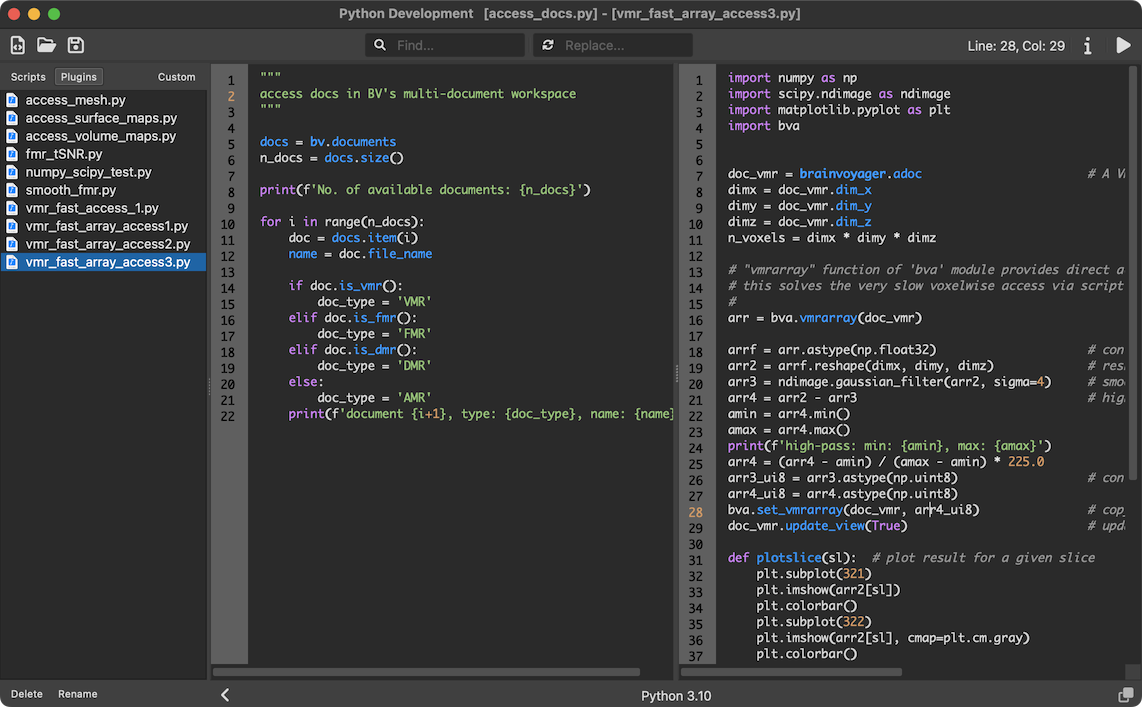
Another highlight of this release for all platforms is the new real-time volume renderer that is partially visible in the screenshot above. While BrainVoyager always had volume rendering capabilities, this is the second time where a new implementation was needed to provide more possibilities and more advanced rendering results. Another major focus of this release was enhanced Python integration: The approach to embed the Python interpreter has been fundamentally changed (based on 'PySide6') to provide more integrated, powerful and future-proof Python support. The latter will allow in a future release to also use BrainVoyager functionality from outside the application enabling the use of any Python development environment. But also the internally offered Python IDE has been substantially improved in this release including, for example, side-by-side editors, search and replace, multi-line indentation and shell commands. A snapshot of the new Python editor is shown below.
BrainVoyager 23 also introduces Lanczos interpolation, which is an optimised windowed sinc interpolation variant that produces at least as good results as standard windowed sinc interpolation but with reduced computation time on CPU and GPU (using Metal on macOS). Furthermore, GIFTI mesh files are now supported as a native format that can be loaded like BrainVoyager .SRF files directly from the Meshes menu; GIFTI functional map ('.func.gii') and label files ('.label.gii') are also supported. These as well as many other new features, enhancements and bug fixes are described in the release notes and the updated 23.0 User's Guide that can be viewed within BrainVoyager, but also here online as well as via a beautiful eBook. To install BrainVoyager 23 for your operating system, go directly to the Downloads page of your platform:
Another highlight of this release for all platforms is the new real-time volume renderer that is partially visible in the screenshot above. While BrainVoyager always had volume rendering capabilities, this is the second time where a new implementation was needed to provide more possibilities and more advanced rendering results. Another major focus of this release was enhanced Python integration: The approach to embed the Python interpreter has been fundamentally changed (based on 'PySide6') to provide more integrated, powerful and future-proof Python support. The latter will allow in a future release to also use BrainVoyager functionality from outside the application enabling the use of any Python development environment. But also the internally offered Python IDE has been substantially improved in this release including, for example, side-by-side editors, search and replace, multi-line indentation and shell commands. A snapshot of the new Python editor is shown below.

BrainVoyager 23 also introduces Lanczos interpolation, which is an optimised windowed sinc interpolation variant that produces at least as good results as standard windowed sinc interpolation but with reduced computation time on CPU and GPU (using Metal on macOS). Furthermore, GIFTI mesh files are now supported as a native format that can be loaded like BrainVoyager .SRF files directly from the Meshes menu; GIFTI functional map ('.func.gii') and label files ('.label.gii') are also supported. These as well as many other new features, enhancements and bug fixes are described in the release notes and the updated 23.0 User's Guide that can be viewed within BrainVoyager, but also here online as well as via a beautiful eBook. To install BrainVoyager 23 for your operating system, go directly to the Downloads page of your platform:
- BrainVoyager 23.0 for Apple Silicon and Intel Macs
- BrainVoyager 23.0 for Windows 10 and 11
- BrainVoyager 23.0 for Linux
Exhibition at Annual OHBM Meeting in Montreal, Canada
18 07, 23
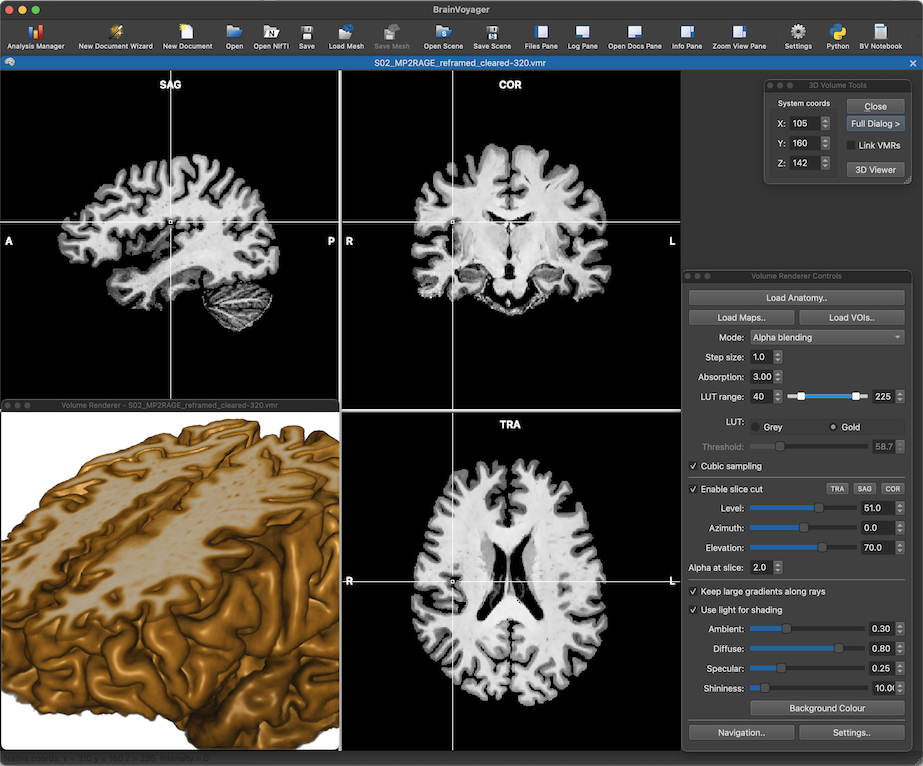
If you are attending this years Human Brain Mapping conference in Montreal, Canada, please visit our booth (#206) and you'll meet BrainVoyager experts answering your questions! We are thrilled to show you several new applications and major updates of existing software, including BrainVoyager 23 with an amazing new volume renderer (see figure below). For the first time, BrainVoyager natively supports Apple Silicon hardware with Metal GPU performance that enables energy-efficient reproducible analyses of large datasets, beautiful real-time renderings and integration of native (arm64) Python as well as accelerated deep learning using TensorFlow. Because of the similarities between M1/M2 iPads and M1/M2 laptop and desktop Macs, desktop-quality BrainVoyager software will also be released for tablets - at our booth we demonstrate, for example, the beta version of BrainViewer for iPad with a touch-driven user interface. Like the recently released Brain Tutor 3D app, BrainViewer not only runs on all major desktop and mobile platforms but it also comes as a WebApp (based on WebAssembly) allowing to launch the app from most web browsers without installing anything. The BrainViewer app not only supports BrainVoyager files but also NIfTI and GIfTI files in a powerful app that can be used to visualise volume and surface datasets with overlaid regions, maps and invoked time courses. The app is so general that it can be used by BrainVoyager users as well as users of other software packages to inspect datasets and to create beautiful publication-ready figures and movies.
We also highlight our latest open source contributions to the software LayNII for state-of-the-art laminar and columnar fMRI analysis. We also present Turbo-BrainVoyager 4.4 that now supports multi-slice DICOM files (used increasingly by SIEMENS) and NIfTI files (used by PHILIPS). TBV 4.4 also comes with a new real-time data simulator that is also included in the Turbo-BrainVoyager 4.4 EDU version. Our EDU products are the ideal tools to teach (e.g. in courses) and (self-)learn how to analyse and visualise structural and functional neuroimaging data from basic to advanced tools including scripting and programming using Python. In our partnership with NIRx (booth #202), we are excited to present the latest version of our real-time fNIRS software Turbo-Satori and a major new version of our fNIRS offline software Satori that now integrates calls into Python tools, for example, in graphically designed data processing workflows.
We also highlight our latest open source contributions to the software LayNII for state-of-the-art laminar and columnar fMRI analysis. We also present Turbo-BrainVoyager 4.4 that now supports multi-slice DICOM files (used increasingly by SIEMENS) and NIfTI files (used by PHILIPS). TBV 4.4 also comes with a new real-time data simulator that is also included in the Turbo-BrainVoyager 4.4 EDU version. Our EDU products are the ideal tools to teach (e.g. in courses) and (self-)learn how to analyse and visualise structural and functional neuroimaging data from basic to advanced tools including scripting and programming using Python. In our partnership with NIRx (booth #202), we are excited to present the latest version of our real-time fNIRS software Turbo-Satori and a major new version of our fNIRS offline software Satori that now integrates calls into Python tools, for example, in graphically designed data processing workflows.

Brain Tutor 3D Web App
11 02, 23
Brain Tutor 3D is now available as a Web App - try it out here! Brain Tutor 3D is the first app that we put on the web, which is made possible by the impressive WebAssembly (Wasm) infrastructure that allows to compile C++ (and other languages) to a cross-platform highly efficient byte code that runs in all major desktop and mobile web browsers. Wasm supports WebGL/OpenGL, which is ideal for BrainVoyager related software that is based on efficient 3D graphics rendering. There are also limitations. As opposed to an installed app, the app is downloaded each time when visiting the page on the web (it is usually kept in memory by the web browser as long as the respective browser page is not closed). While the whole software is packaged in a small wasm file of about 60 MB, it will take a few seconds until the file is downloaded and the web app is running. If one uses an app more than a few times, it may be a better option to download the corresponding desktop or mobile app. Wasm apps are also 32-bit programs (the desktop and mobile versions are 64-bit apps) posing restrictions on handling very large datasets. Wasm apps can be employed easily on the Web, which has the advantage that it runs by clicking on a link instead of downloading and installing different applications compiled for specific operating systems. On the developer side, web employment is easy avoiding creation of platform-specific installers as downloadable packages or as apps for App Stores that come with many extra requirements. Despite avoiding approval from App Stores, WebAssembly apps are secure to use since they run in a sandbox of the browser. This makes it more challenging to use files from the local file system for processing or visualization. This is, however, possible as long as the user explicitly selects (and thereby approves) files from the local system for uploading into the sandbox. This will allow our next web app, Brain Viewer, to process your custom BrainVoyager files but also NIfTI, GIfTI and other files in a powerful web application - stay tuned!

Exhibition at Annual OHBM Meeting in Glasgow
19 06, 22
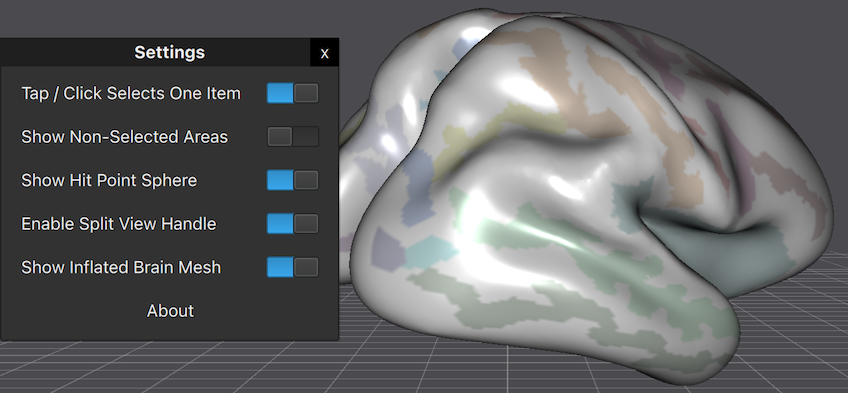
If you are attending this years in-person Human Brain Mapping conference in Glasgow, please visit our booth (#13) and you'll meet BrainVoyager experts answering your questions! We are thrilled to show you several new applications and major updates of existing software, including BrainVoyager 23 with an amazing new volume renderer and, for the first time, a real-time path tracer producing stunning visualizations. BrainVoyager 23 fully supports Apple Silicon hardware with Metal GPU performance that enables efficient reproducible data analyses, beautiful real-time renderings and integration of native (arm64) Python as well as accelerated deep learning using both TensorFlow and PyTorch. Because of the similarities between M1/M2 iPads and M1/M2 laptop and desktop Macs, desktop-quality BrainVoyager software will also soon be released for iPad - at our booth we demonstrate, for example, BrainVoyager Viewer for iPad with a touch-driven user interface.
We also highlight our latest open source contributions to the software LayNII for next generation laminar fMRI analysis. We also demonstrate advances of Turbo-BrainVoyager such as support for NIfTI files and a new real-time data simulator that is also included in the new Turbo-BrainVoyager EDU product. Our EDU products are the ideal tools to teach (e.g. in courses) and (self-)learn how to analyse and visualise structural and functional neuroimaging data from basic to advanced tools including scripting and programming using Python. In our partnership with NIRx, we are excited to present the latest version of our real-time fNIRS software Turbo-Satori and the new software Satori for multi-session, multi-subject fNIRS group analyses featuring advanced analysis and visualisation options. In our partnership with Mag & More, we are excited to show the latest features of the new PowerMAGView! 2.0 release.
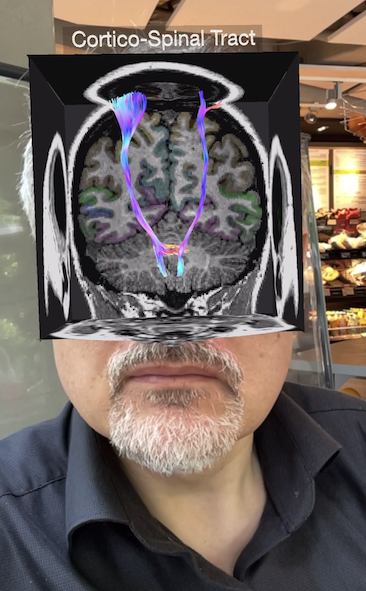
If you like augmented reality apps, try out the Scan My Brain app, which is fun to use and teaches knowledge about the brain (see screenshot below from the developer); you can also try it out yourself on your iPhone (or iPad) - just get the app from the AppStore using the download link of the public beta version:
https://testflight.apple.com/join/SwsYY3uS
If not yet installed, you will be asked to first load the 'TestFlight' app, which is Apple's tool to distribute beta versions.
We also highlight our latest open source contributions to the software LayNII for next generation laminar fMRI analysis. We also demonstrate advances of Turbo-BrainVoyager such as support for NIfTI files and a new real-time data simulator that is also included in the new Turbo-BrainVoyager EDU product. Our EDU products are the ideal tools to teach (e.g. in courses) and (self-)learn how to analyse and visualise structural and functional neuroimaging data from basic to advanced tools including scripting and programming using Python. In our partnership with NIRx, we are excited to present the latest version of our real-time fNIRS software Turbo-Satori and the new software Satori for multi-session, multi-subject fNIRS group analyses featuring advanced analysis and visualisation options. In our partnership with Mag & More, we are excited to show the latest features of the new PowerMAGView! 2.0 release.
If you like augmented reality apps, try out the Scan My Brain app, which is fun to use and teaches knowledge about the brain (see screenshot below from the developer); you can also try it out yourself on your iPhone (or iPad) - just get the app from the AppStore using the download link of the public beta version:
https://testflight.apple.com/join/SwsYY3uS
If not yet installed, you will be asked to first load the 'TestFlight' app, which is Apple's tool to distribute beta versions.